

The flowering plant tree of life, much like our own family tree,?allows us to understand how different species are related to each other. The tree of life is?revealed?by comparing DNA sequences between different species to identify changes (mutations) that accumulate over time like a molecular fossil record. Our understanding of the tree of life is rapidly improving due to advances in DNA sequencing technology.
A vast DNA tree of life brings open access DNA sequences of >9500 flowering plants was recently achieved by?scientists?from the Royal Botanic Gardens, Kew, together with collaborators from the?Kunming Institute of Botany, Chinese Academy of Sciences?and?around the globe, This invaluable resource lets us answer key questions about modern plant life and look back in time to its origins. Their study was recently published in?Nature.
?
A key advantage of the approach is that it can be used to sequence a wide?range?of plant material, old and new, even when the DNA is badly damaged. The vast treasure troves of dried plant material in the world’s herbarium collections, which?include?nearly 400 million scientific?plant?specimens, can now be studied genetically.
Using such specimens, the?researchers?sequenced a sandwort specimen (Arenaria globiflora) collected nearly 200 years ago in Nepal and, despite the poor quality of its DNA, were able to place it in the tree of life. They?even analyzed extinct plants, such as the Guadalupe Island olive (Hesperelaea palmeri), which has not been seen alive since 1875. In fact, 511 of the species sequenced are already?threatened?with?extinction, according to the International Union for Conservation of Nature Red List, including three more like?Hesperelaea?that are already extinct.
Of the?9,506 species sequenced for this study, over 3,400?were derived?from material sourced from 163 herbaria in 48 countries,?with?additional material from plant collections around the world (e.g., DNA banks, seeds, and living collections). Among the species sequenced, more than 800 had never had their DNA sequenced before.??This sequencing was essential to fill in important knowledge gaps and shed new light on the evolutionary history of flowering plants.?The?researchers?also benefited from publicly available data for?more than?1,900 species, highlighting?the?value of the open science approach?to?future genomic research.
Despite the contrasting biological properties of the nuclear and plastid genomes (e.g.,?size, copy number, mode of inheritance, recombination, and evolutionary rate),?which can lead to conflicting phylogenic trees,?the results?largely support the mostly plastid-based phylogenetic classification of the Angiosperm Phylogeny Group?IV. For example, 58 of the 64 currently accepted orders and 406 of the 416 families were recovered as monophyletic (excluding artifacts). The most striking exception is the non-monophyly of Asteraceae, the largest angiosperm family, which?includes?sunflowers and their relatives.?The?tree produced by this study also confirms 85% of the relationships among families recovered by the phylogenomic angiosperm tree?using plastomes by scientists from?the?KIB.
Flowering plants originated over 140 million years ago,?after which they rapidly overtook other vascular plants. Darwin was?puzzled?by the seemingly sudden appearance of such diversity in the fossil record?and?wrote:?“The rapid development,?as far as we can judge,?of all the higher plants within recent geological times is an abominable mystery.”
Using?200 fossils, the?researchers traced their tree of life?back in?time?to show?how flowering plants evolved?over?geological time. They found that early flowering plants did indeed explode in diversity, as Darwin noted. The rapid development of these plants gave rise, shortly after their origin, to over 80% of the major lineages that exist today. However, this trend then declined to?a more stable?rate for the next 100 million years,?until another surge in diversification?occurred?about 40 million years ago, coinciding with a global drop?in temperature.
These new?findings would have fascinated Darwin and will surely help today’s scientists?as they?grapple?with the challenges of understanding how and why species diversify.
The flowering plant tree of life has enormous potential?for?biodiversity research. This is because, just as one can predict the properties of an element based on its position in the periodic table, the location of a species in the tree of life allows us to predict its properties. The new data will therefore?be invaluable?in improving?many areas of science and beyond.
To?make?this?possible, the tree and all?its underlying?data have been made openly and freely available?to both the public and?the?scientific community, including through the?Kew Tree of Life Explorer. The?researchers believe?that?such open access is key to democratizing access to scientific data around?the?world.
Open access will also help scientists make the?most?of the data, such as combining it with artificial intelligence to predict which plant species may?contain?molecules with medicinal potential. Similarly, the tree of life can be used to better understand and predict how pests and diseases?will?affect plants in the future. Ultimately, the?researchers?note, the applications of this data will be driven by the ingenuity of the scientists?who?access it.
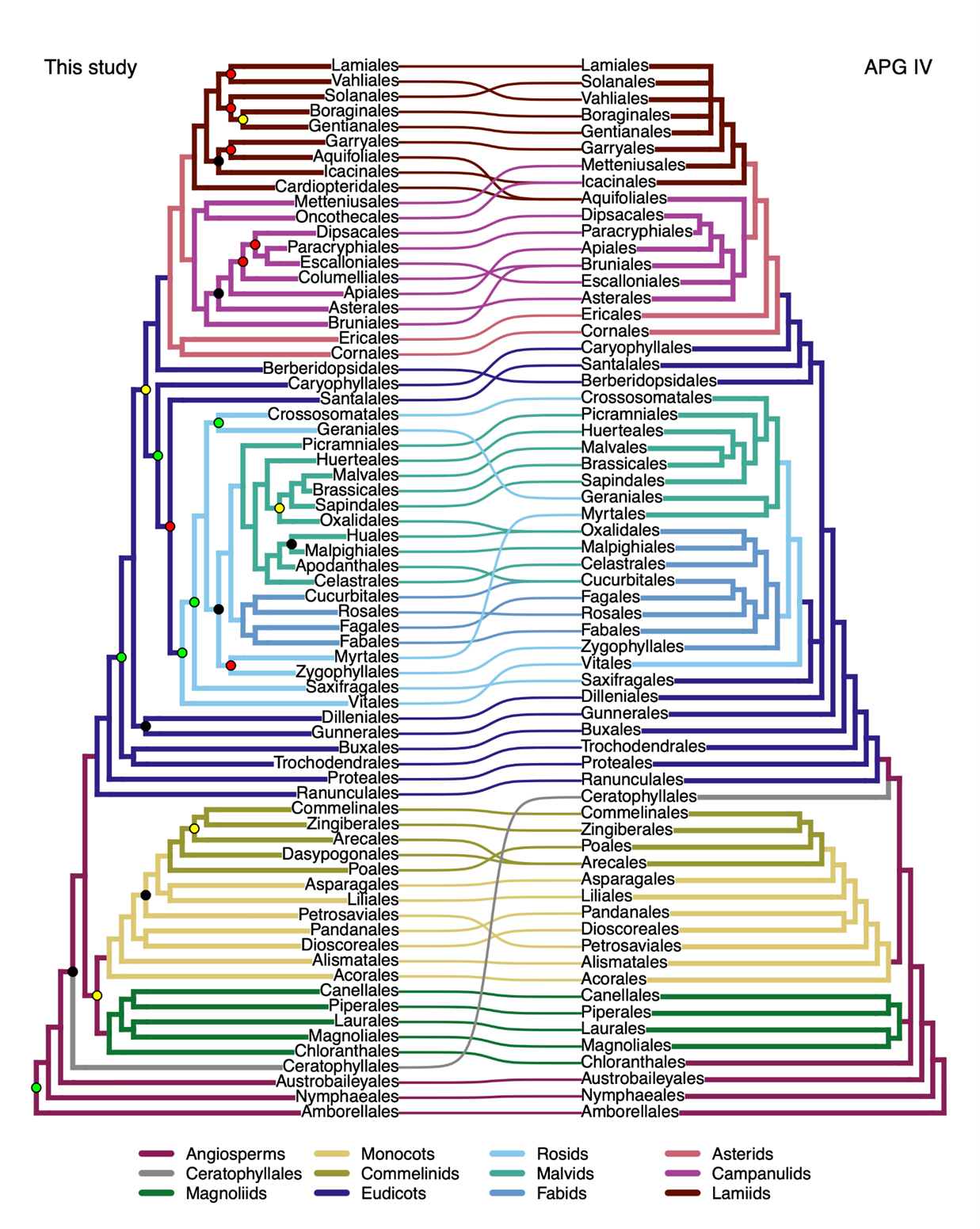
Fig. 1? Tanglegram at ordinal level between this work (left) and the APG IV schematic
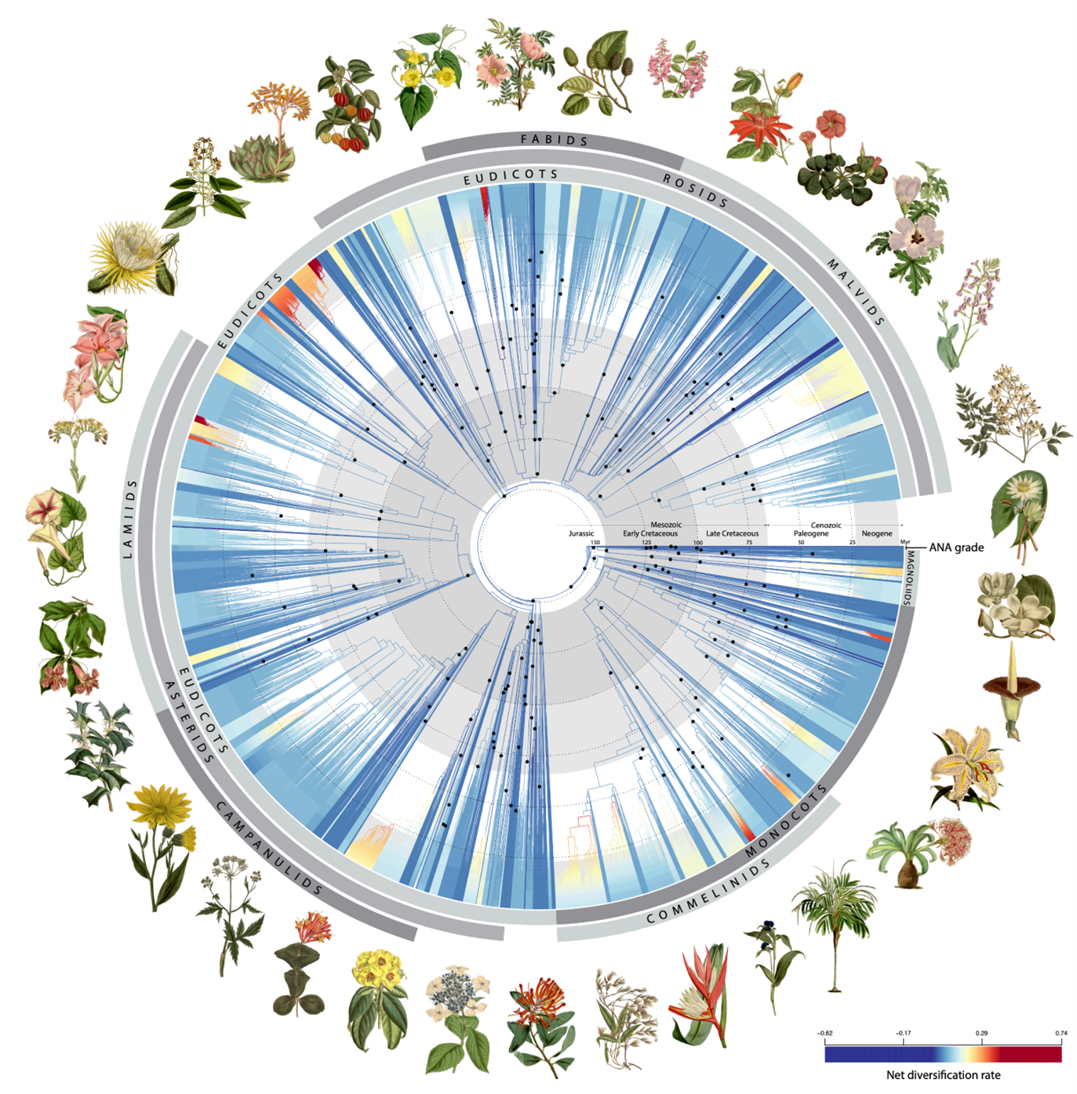
Fig. 2? Time-calibrated phylogenetic tree for angiosperms based on Angiosperms353 nuclear gene datasets.